All published articles of this journal are available on ScienceDirect.
Polymorphisms in X-Ray Repair Cross-Complementing Group 1 Gene: Haplotypes, Breast Cancer Risk and Individual Radiosensitivity
Abstract
The aim of this paper is to analyse the role exerted by X-ray repair cross-complementing group 1 (XRCC1) genetic polymorphisms and haplotypes in increasing breast cancer risk and in modulating radiotherapy-induced adverse reactions. An Italian cohort of breast cancer patients and a matching group of healthy controls were genotyped for XRCC1-77T>C, Arg194Trp and Arg399Gln polymorphisms. Our data indicated that polymorphisms at codon 399 and at -77 position of the 5’-untraslated region both contribute to cancer risk. We also showed that the haplotype H3, containing the wild-type allele at codon 194 and the variant alleles at codon 399 and at -77 position is significantly associated with an increased risk of breast cancer. We found no statistical association between XRCC1 SNPs and individual radiosensitivity.
INTRODUCTION
Breast cancer (BC) is the most common cause of cancer death in women worldwide and radiotherapy immediately after surgery is the standard treatment for this disease. In the recent years, many efforts have been devoted to identify molecular factors that could increase both the risk of disease development and the induction of adverse effects following radiation therapy. Among several factors, our group and many others have focused on polymorphisms of genes involved in DNA damage repair, in particular XRCC1 (X-ray repair cross-complementing group I) gene [1-4]. XRCC1 plays a key role in base excision repair and is also involved in other DNA repair pathways as single-strand break repair and non-homologous end joining [5]. In general, single-nucleotide polymorphisms (SNPs) in this gene may affect DNA repair function. It has been demonstrated that some of the most studied variants in XRCC1 coding region alter protein structure, while SNPs in the upstream regulatory sequence affect gene expression, therefore possibly modu-lating the response to environmental mutagens/carcinogens and cancer risk [6].
Most recently, genome-wide association studies (GWAS) have identified a great number of low-penetrance variants associated with BC susceptibility, some of which are known to influence the development of different cancers [7, 8]. Several studies on the association between SNPs and radiotherapy toxicity have been published, most of them based on a candidate gene approach. Anyway, up to now, there are not conclusive data on this field [9, 10].
The aims of our work were to confirm our previous data suggesting the contribution of XRCC1 genetic polymorphisms in increasing BC risk and to evaluate the potential role of these variants in radiotherapy-induced adverse reactions. The present retrospective study was performed on 92 breast cancer patients, selected for the homogeneity in total dose applied and the follow up time and 104 control subjects matched for age and lifestyle. In particular, attention was devoted to three genetic variants of XRCC1 gene: -77T>C substitution (rs3213245), located in the 5’-untranslated region, C>T substitution in codon 194 (rs1799782) and G>A substitution in codon 399 (rs25487), which result in non-conservative amino acid changes at highly conserved regions (respectively Arg194Trp and Arg399Gln). We also considered the eight haplotypes generated by the combination of these three loci.
MATERIALS AND METHODS
Study Subjects
Ninety-two breast cancer patients and one hundred-four controls were involved in this study. Patients aged between 40 and 85 years (mean age = 63 ± 7.5) were recruited at Radiotherapy Unit of S. Camillo-Forlanini Hospital and Radiation Oncology Unit of S. Pietro Hospital (Rome, Italy), after breast conserving surgery and before receiving primary radiotherapy. Forty-three of these patients were recruited from May 2007 to May 2008 and were already part of a previous study [1], while the remaining were recruited from May 2011 to May 2012. Patients with family histories of breast cancer and patients treated with a mixed regimen (radiotherapy in combination with chemotherapy) were not included in the study.
Patients were treated in both Radiotherapy Units with a standard regimen of radiation therapy consisting of a reference dose of 50 Gy given with 2 Gy per fraction, five times per week, to the whole breast, irradiated by a 6 MV photon beam and with a boost irradiation (10 Gy) to the tumour bed administered with 9-12 MeV electrons. Both early and late effects were evaluated by experienced radiooncologists according to Radiation Morbidity Scoring Scheme (EORTC/RTOG), graded on 5-point ordinal scales (0 meaning absence of radiation effect and 5 the effects leading to death), with high-grade toxicity considered as grade > 2 [11].
The following clinical radiation skin reactions within the radiation field of the breast were documented during treatment: erithema, desquamation, decreased sweating and edema. Common late radiation effects on the skin and subcutaneous tissues (i.e., effects that first occur 90 days or more after initiation of RT) include fibrosis, telangiectasia and atrophy.
Early side effects were documented in 92 patients during four steps: 1) before the beginning of radiotherapy; 2) at a cumulative dose of 36–42 Gy; 3) at the end of radiotherapy (about 60 Gy cumulative dose) and 1 month later. Late side effects were documented in 45 patients and the median follow-up time was 52 months.
The study was approved by the Ethical Committee of the participating hospitals and written informed consents were obtained from the study subjects.
Healthy volunteers aged between 40 and 82 (mean age 58 ± 7.4) were selected as a matching group. Subjects with a prior history of oncologic diseases were excluded. All donors completed a written questionnaire to obtain information related to their lifestyle and medical history. Each study subjects contributed to the study with a single blood draw.
Genotyping
Genomic DNA was isolated from 200 µl of whole blood, using the EURx GeneMATRIX Quick Blood DNA Purification Kit (http://www.eurx.com.pl). The RFLP–PCRs for genotyping rs25487 (XRCC1-399), rs1799782 (XRCC1-194) and rs3213245 (XRCC1 -77T>C) variants were performed as previously described [12-14], PCR products were digested with specific restriction enzymes that recognized and cut either the wild-type or variant sequence site.
Statistical Analysis
Statistical analysis of the data was performed using the GraphPad Prism 5 software (http://www.graphpad.com). We used the χ2 test to verify the Hardy-Weinberg equilibrium of XRCC1 SNPs and to calculate the association between XRCC1 haplotypes and radiotherapy adverse reactions. Fisher’s exact test was used for the comparison of 2 x 2 contingency tables. The odds ratios (ORs) and 95% confidence intervals (CIs) were calculated to assess BC risk and risk of developing adverse reactions after radiotherapy. We considered p dered p ≤ 0.05 to be si 0.05 to be significant for all analyses.
RESULTS
We investigated three XRCC1 variant alleles at -77 position, at codons 194 and 399. The genotype distributions were in Hardy-Weinberg equilibrium (p ≥ 0.5 at χ2 test) except for -77T>C SNP in the group of controls. We found that the allele frequencies for the studied polymorphisms in the controls were similar to those previously reported in Caucasian control subjects [15, 16], namely 0.34 for the variant C allele at position -77, 0.07 for the variant T allele at codon 194 and 0.31 for the variant A allele at codon 399. As for the XRCC1-399 variant allele, we found a significantly higher frequency (0.41) in BC patients compared to controls (p < 0.05). In Table 1, we show the associations between the three XRCC1 polymorphisms, XRCC1 haplotypes and BC risk. We found a positive association between XRCC1-77T>C SNP and BC occurrence. This association is significant (p = 0.03) considering the -77 heterozygous (T/C) genotypes. When the sum of the heterozygous and the homozygous genotypes (T/C+C/C) was considered, the p value was close to significance (p = 0.06). A more clear positive association was found between XRCC1-399 SNP and BC occurrence. This association was significant both for heterozygous (p = 0.05) genotypes and when considering the sum of the heterozygous and the homozygous genotypes (p = 0.03).
Association between XRCC1 SNPs, XRCC1 haplotypes and BC risk.
| SNPs | Cases (%) N = 92 |
Controls (%) N = 104 |
OR (95% CI) |
|---|---|---|---|
| XRCC1 -77 T>C | |||
| T/T | 35 (38.0) | 52 (50.0) | 1.00 (Ref.) |
| T/C | 42 (45.7) | 33 (31.7) | 1.89 (1.01 - 3.54)** |
| C/C | 15 (16.3) | 19 (18.3) | 1.17 (0.53 - 2.61) |
| T/C+C/C | 57 (62.0) | 52 (50.0) | 1.63 (0.92 - 2.88) |
| XRCC1 – 194 C>T | |||
| C/C | 79 (85.9) | 91 (87.5) | 1.00 (Ref.) |
| C/T | 13 (14.1) | 12 (11.5) | 1.25 (0.54 - 2.89) |
| T/T | 0 (0) | 1 (1.0) | NC |
| C/T+T/T | 13 (14.1) | 13 (12.5) | 1.15 (0.50 - 2.63) |
| XRCC1 – 399 G>A | |||
| G/G | 33 (35-9) | 52 (50.0) | 1.00 (Ref.) |
| G/A | 43 (46.7) | 39 (37.5) | 1.74 (0.94 - 3.21)* |
| A/A | 16 (17.4) | 13 (12.5) | 1.94 (0.83 - 4.55) |
| G/A+A/A | 59 (64.1) | 52 (50.0) | 1.79 (1.01 - 3.17)** |
| Haplotypesa | |||
| H1 C-C-G | 24 (26.1) | 34 (32.7) | 1.00 (Ref.) |
| H2 T-C-A | 26 (28.3) | 33 (31.7) | 1.12 (0.54-2.32) |
| H3 C-C-A | 27 (29.3) | 15 (14.4) | 2.55 (1.12-5.79)*** |
| H4 T-C-G | 2 (2.2) | 9 (8.7) | 0.31 (0.06-1.59) |
| H5 T-T-G | 1 (1.1) | 7 (6.7) | 0.20 (0.02-1.76) |
| H6 T-T-A | 6 (6.5) | 3 (2.9) | 2.83 (0.64-12.47) |
| H7 C-T-G | 6 (6.5) | 2 (1.9) | 4.25 (0.79-22.89) |
| H8 C-T-A | 0 (0) | 1 (1) | NC |
Abbreviations: NC, not calculated; Ref., reference genotype or haplotype.
a The haplotype is defined as the combination of alleles present at position -77(T>C), codons 194(C>T) and 399(G>A), respectively.
Fisher’s test *p = 0.05; **p = 0.03; *** p = 0.02.
Association between XRCC1 SNPs and early and late adverse reactions to radiotherapy.
| Genotype | Early Effects Cases N=92 |
OR (95% CI) | Late Effects Cases N=45 |
OR (95% CI) | ||
|---|---|---|---|---|---|---|
| G>2 | G<2 | G>2 | G<2 | |||
| XRCC -77T>C | ||||||
| T/T | 8 | 27 | 1.00 (Ref.) | 3 | 11 | 1.00 (Ref.) |
| T/C | 15 | 27 | 1.87 (0.68 – 5.15) | 3 | 17 | 0.65 (0.11 - 3.80) |
| C/C | 5 | 10 | 1.69 (0.45 -6.40) | 1 | 10 | 0.37 (0.03 – 4.12 |
| T/C+C/C | 20 | 37 | 1.82 (0.70 -4.76) | 4 | 27 | 0.54 (0.10 – 2.84) |
| XRCC1-194C>T | ||||||
| C/C | 23 | 56 | 1.00 (Ref.) | 5 | 35 | 1.00 (Ref.) |
| C/T | 4 | 9 | 1.08 (0.30 -3.87) | 2 | 3 | 4.67 (0.62 – 35.19) |
| XRCC1-399G>A | ||||||
| G/G | 13 | 20 | 1.00 (Ref.) | 3 | 21 | 1.00 (Ref.) |
| G/A | 11 | 32 | 0.53 (0.20 -1.41) | 3 | 13 | 1.61 (0.28 – 9.24) |
| A/A | 4 | 12 | 0.51 (0.14 -1.94) | 1 | 4 | 1.75 (0.14 – 21.40) |
| G/A+A/A | 15 | 44 | 0.52 (0.21 – 1.30) | 4 | 17 | 1.65 (0.32 – 8.39) |
Abbreviations: NC, not calculated; Ref., reference genotype or haplotype.
We obtained eight different XRCC1 haplotypes from the combination of the three SNPs studied in our population (Table 1). In the controls, the most frequently found haplotype was H1 (32.7%), containing the C variant at position -77 together with the wild-type alleles at codons 194 and 399 and it was considered by us as reference. The haplotype H3, containing the wild-type allele at codon 194 and the variant alleles at codon 399 and at -77 position was found in the 29.3% of the cases and in the 14.4% of the controls and was significantly (p < 0.02) associated with an increased risk of BC.
We also evaluated in BC patients a possible association between XRCC1 gene variants and the risk of developing radiotherapy-induced severe acute and late skin reactions on normal tissue.
As regards the acute side effects, 28 cases out of 92 (30.4%) suffered from severe toxicity (Gity (G≥2), w2), while 7 patients out of 45 (15.6%) showed severe late adverse reactions (Table 2). None of the three XRCC1 SNPs showed a significant association with radiation sensitivity, as regards the onset of either early or late side effects.
In Fig. (1), we reported the distribution of XRCC1 H1, H2 and H3 haplotypes in BC patients without severe toxicity (G<2) and in patients with G≥2, respectively related to early and late effects. As regards the early effects, the most represented haplotypes in patients with G<2 are H2 and H3 (31.2% each), while the most common haplotype in the G≥2 group is H1 (35.7%). Furthermore if we consider the distribution of patients based on the reactions within the single haplotype, we found the larger number of G≥2 patients in the H1 haplotype (10 of 24 vs 6 of 26 in H2 or 7 of 27 in H3).
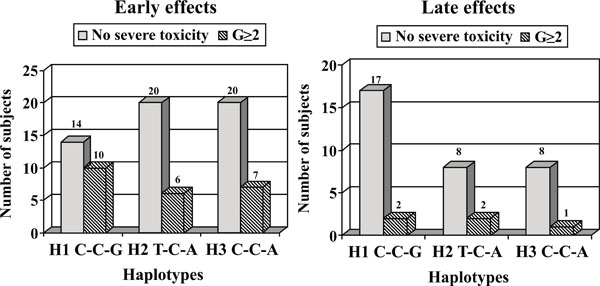
Distribution of XRCC1 H1, H2 and H3 haplotypes in breast cancer patients without severe toxicity (G<2) and in patients with grade >2, respectively related to early and late effects.
As concerning late normal tissue complications, the most commonly represented haplotype in patients without severe toxicity is H1 (44.7%) as well as in patients with G≥2, at the same rate of H2 (28.6 % each). However, considering the distribution of patients within the single haplotype, we found that G≥2 patients mostly present H2 haplotype (2 of 10 vs 2 of 19 for H1).
DISCUSSION
We previously reported a strong significant association between breast cancer occurrence and the presence of the XRCC1 variant allele (399-Gln) in a cohort of Italian patients [17]. In the present study we examined a larger number of BC patients with the aim to assess whether SNPs and haplotypes in XRCC1 gene were associated with an increased risk of both developing the disease and adverse reactions after radiotherapy.
Results showed that XRCC1-399 SNP significantly increased susceptibility to breast cancer, confirming our previous data obtained on a smaller sample of 43 BC patients [1, 17] and suggests a dominant inheritance model for the XRCC1-399 variant allele.
The results of association studies between XRCC1-399 SNP and BC risk have been summarized in several meta-analyses [18-22], nevertheless yielding conflicting results. Huang and co-workers [18] showed that Arg399Gln SNP is associated with a trend of increased breast cancer risk when using both dominant and recessive models, even if it is weakly related with breast cancer in Caucasians. In a more recent meta-analysis, Bu et al. [19] observed a significant association between XRCC1 Arg399Gln variant and risk of breast cancer in the American population, considering both the dominant and the additive model, but they did not find any association in the recessive model. On the contrary, other studies suggested no association in any inheritance models in Caucasian population [20-22].
Our data on the association between XRCC1-77T>C SNP and BC risk should be considered with caution since statistical significance was achieved only considering heterozygous genotypes. Studies investigating the association of XRCC1-77T>C SNP and cancer risk have reported conflicting results. A meta-analysis including 13 studies showed a significant association between the C variant of XRCC1-77T>C polymorphism. In the subgroup analysis based on cancer type, the XRCC1-77C variant was significantly associated with the risk of breast cancer and lung cancer, but in the subgroup analysis based on ethnicity, this association was still significant in the Asian population, but not in Caucasians [23].
We also determined the association between XRCC1 haplotypes, based on the three SNPs investigated in our study, and susceptibility to breast cancer.
The haplotype H3 (wild-type allele at codon 194 and variant alleles at codon 399 and at -77 position) showed a positive association with an increased risk of BC, in agreement with our previously published paper [1] and in contrast with the results of a meta-analysis of case–control studies conducted by Saadat and co-workers [24]. So far, only a few XRCC1 haplotypes analyses have included the SNP in the promoter [16, 4]. Our data are partially in agreement with the only XRCC1 haplotype analysis performed on a Caucasian population, as previously discussed [1] and in contrast with a more recent study on a Chinese population, where the significant increase in BC risk was observed for the haplotype containing the variant allele at position -77 and the wild-type alleles at codons 194, 280 and 399 [4].
Our overall results showed that XRCC1 SNPs and haplotypes may contribute to the genetic risk for BC. Although associations between SNPs in XRCC1 and other DNA repair genes and susceptibility to breast cancer has been widely investigated, clearly defined results in this context have not yet been achieved. In particular, Arg399Gln SNP, the most common variant in the XRCC1 gene, is an important polymorphism related to sporadic breast cancer susceptibility. Nevertheless, literature data showed a weak association of this SNP with BC risk, that is stronger only within some ethnicities [25]. In general, the effect of SNPs on the occurrence of breast cancer and other tumour types is usually only slightly statistically significant.
In some studies, SNP-SNP interactions have been examined to evaluate epistatic effects contributing to BC. Specific SNP pairs, selected within genes in DNA repair pathways or in other DNA metabolism pathways, showed a statistical association with BC risk [26]. Significant trends in BC risk were also observed in association with an increasing number of risk alleles in different DNA repair genes [27]. Therefore, association studies on haplotypes of genes as XRCC1 and on the interaction between SNPs in DNA repair genes should be encouraged because although a single SNP may have a negligible effect, interactions between variants in different genes could significantly affect cancer risk.
In order to implement the use of SNPs in the practice of clinical medicine, the future challenges will be not only identifying causative low-penetrance variants but also determining how these SNPs can interact with each other and with environmental and pathobiological factors. Furthermore, since breast cancer is a highly heterogeneous disease, morphological and immunohistochemical characteristics of the tumour are currently used in the prognosis and in treatment planning. Therefore the possible association of SNPs in XRCC1 and other DNA repair genes with histological type of tumour and hormone receptor status is an interesting topic to develop and is the core of our current studies on breast cancer patients.
Concerning the possible effect of XRCC1 SNPs on radiation toxicity our data, showing no significant association regarding the onset of either early or late side effects, confirm our previous results obtained in a more limited number of patients [1]. In a recent meta-analysis [28], the predictive value of XRCC1 SNPs for side effects in patients undergoing whole breast radiotherapy has been determined. 11 studies mainly comprising Caucasian patients, addressing both acute and late toxicity, were included in the analysis. As we found in our study, no significant association with the XRCC1-399 variant allele was observed. Similar negative results were also found for XRCC1-194 and -77 SNPs. Nevertheless, a predictive value of XRCC1-399 SNP was found in studies with mixed treatment regimens when studies on only late toxicity were excluded. Another meta-analysis including 14 case-control studies, 13 of which were performed in European countries, evaluated the association between XRCC1-399 SNP and the risk of normal tissue injury after radiotherapy in BC patients [29]. This meta-analysis suggests that XRCC1-399 SNP was significantly associated with increased risk of adverse normal tissue reactions after radiation therapy. Since the data published so far are still conflicting, further well-designed studies are needed to clarify the role of XRCC1 polymorphisms in modulating radiotherapy-induced adverse effects in breast cancer patients.
As regards the correlation between XRCC1 haplotypes and radiotherapy side effects, when we considered the distribution of patients according to the severity of early reactions within the single haplotype (Fig. 1), we found a greater number of G>2 patients in the H1 haplotype (41.6% vs about 23% in H2 or H3 haplotypes). This result suggests that the presence of SNP at position -77 could influence the development of acute adverse side effects even if no statistical significance was reached.
When we considered the distribution of patients within the single haplotype according to the severity of late reactions illustrated in Fig. (1), we found that G>2 patients mostly presented with a H2 haplotype (20% vs 10% for H1). This result suggests that the presence of a variant allele at codon 399 could influence the development of late toxicity even if no statistical significance was reached. Even if this finding must be carefully considered due to the small number of subjects, it should be in agreement with our previous hypothesis suggesting that the variant allele in position 399 promotes a faster resolution of DNA damage induced by irradiation. These possibly misrepaired lesions might in part cause genetic instability and consequently apoptotic cell death [12].
To our knowledge, there are no other data on the association between haplotypes, even in other genes besides XRCC1, and normal tissue adverse reactions illustrated in Fig.(1). An interesting and comprehensive meta-analysis conducted by Andreassen has recently shown controversial results on the association between single SNPs and normal tissue complication risk [30]. The first GWAS have identified genetic variants associated with radiotheraphy toxicity, but there isn’t any confirmed SNP so far [10, 31]. Radiogenomics is a promising field in oncology and recent studies are emphasizing the importance of understanding the molecular pathways and genetic components responsible for individual radiosensitivity [32]. One important goal of radiogenomics is to develop SNP-based assays to estimate the risk for an oncologic patient to suffer from radiotherapy-induced adverse reactions. These predictive assays could be used to customize the radiotherapy protocols for both sensitive and resistant patients. In the last years, normal tissue complication probability (NTCP) models, combining genotyping profiles, clinical data and treatment parameters, have been introduced into clinical practice in order to identify patients at risk for developing radiotherapy-induced toxicities. It has been demonstrated that the predictive ability of NTCP models can be significantly improved by incorporating SNP information [33]. Further studies on radiogenomics can contribute to achieve increasingly better individualized radiotherapy protocols and could provide new therapeutic targets in a future perspective of personalized medicine in oncology care.
In this context our study, although having several limitations such as the moderate sample size of cases and controls, has to be considered a contribution to a more exhaustive collection of data.
CONFLICT OF INTEREST
The authors confirm that this article content has no conflict of interest.
ACKNOWLEDGEMENTS
We would like to thank Dr. Daniela Giammarino and Dr. Vittorio Donato from S. Camillo-Forlanini Hospital, Radiation Oncology Unit, Rome, Italy for their important clinical support.
We are indebted to the personnel of the Medicine Service of ENEA Casaccia for their collaboration in selecting and collecting control blood samples.
We are grateful to all patients and controls who helped to make our study possible.


